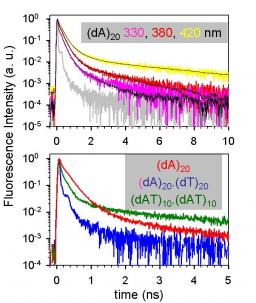
Absorption of ultraviolet light by DNA is known to lead to carcinogenic mutations, but the processes between photon absorption and the photochemical reactions are poorly understood. In their study of the excited-stated dynamics of model DNA helices using femtosecond transient absorption spectroscopy1, Crespo-Hernández et al. observe that the picosecond component of the transient signals recorded for the adenine–thymine oligonucleotide (dA)18•(dT)18 is close to that for (dA)18, but quite different from that for (dAdT)9•(dAdT)9; from this observation, they conclude that excimer formation limits excitation energy to one strand at a time. Here we use timeresolved fluorescence spectroscopy to probe the excited-state dynamics, which reveals the complexity of these systems and indicates that the interpretation of Crespo-Hernández et al. is an oversimplification. We also comment on the pertinence of separating base stacking and base pairing in excited-state dynamics of double helices and question the authors’ assignment of the long-lived signal component found for (dA)18•(dT)18 to adenine excimers. Figure 1a shows the fluorescence decays of (dA)20 at three different wavelengths. Combining these results with our previous measurements obtained for (dA)20 by femtosecond fluorescence upconversion2, at least five exponentials are needed to fit the decays over the 100 femtoseconds to 20 nanoseconds time range. A crucial point is that all time constants vary strongly with the emission wavelength. The same effect is encountered for (dAdT)10•(dAdT)10 and has been reported previously3 for poly(dA)•poly(dT).
•  Physique et chimie pour le vivant et l’environnement › Physique et vivant / Physics and life
Physique et chimie pour le vivant et l’environnement › Physique et vivant / Physics and life